2026-03-10 Qiagen RNeasy Extraction Tests, Cryosectioned tissue optimization
- Extracting RNA from PAXgene fixed tissue at various stages of decalcification
- 3/16/26: All final blocks embedded, details in google sheet and notebook
- Embedded blocks were sectioned 3/17 and 3/19:
- 3/25/26: Tissue scrolls extraction
Extracting RNA from PAXgene fixed tissue at various stages of decalcification
Timeline
- 3/10/26: Corals fixed
- 3/11/26: Fixative replaced with stabilizer
- 3/12 - 3/14: 48 hours of decalc, MON and 3/4 POC embedded
- 3/12 - 3/15: 72 hours of decalc for 2 POR + 1 POC
- 3/12 - 3/16: 96 hours of decalc for 3 POR
- 3/17 & 3/19: Sectioning
- 3/20: All slides H&E Stained
- 3/20 + 3/25: Imaging with EVOS7000
- 3/25: Extraction of 6 scrolls
Protocol: Qiagen RNeasy Kit
Continued testing from first test of this kit for RNA QC here and here
Notes:
- β-Mercaptoethanol (β-ME) must be added to Buffer RLT before use. Add 10 µl β-ME per 1 ml Buffer RLT. Dispense in a fume hood and wear appropriate protective clothing.
- Buffer RPE is supplied as a concentrate. Before using for the first time, add 4 volumes of ethanol (96–100%) as indicated on the bottle to obtain a working solution.
Procedure:
- Extraction protocol was modified from the standard RNeasy protocol for paxgene-fixed tissue sections in OCT
- Modifications based on the protocol of the Qiagen PAXgene Tissue RNA/miRNA Kit (handbook and cryo-embedding specific modifications)
- Make sure all buffers are prepared appropriately
- Label tubes + clean bench
- Tissue prep steps (tissue scroll extraction protocol)
- Tubes were taken out of freezer on dry ice and 150 uL RLT (w/ 1% BME - 1000 + 10 uL) was added and the scrolls were dissolved and mixed with a wide-bore tip
- Then, 290 μl RNase-free water and 10 μl Proteinase K were added and briefly mixed by vortex
- Incubate the tissue on a shaker–incubator for 15 min at 56°C and 1400 rpm
- Centrifuge for 3 min at maximum speed (but do not exceed 20,000 x g).
- Carefully remove 350 uL of the supernatant by pipetting, and transfer it to a new 1.5 mL tube
- Add 1 volume of 70% ethanol* to the cleared lysate, and mix immediately by pipetting.
- Transfer up to 700 µl of the sample to an RNeasy spin column placed in a 2 ml collection tube (supplied).
- Close the lid gently, and centrifuge for 30 s at 15,000 rcf. Discard the flow-through.
- If the sample volume exceeds 700 µl, centrifuge successive aliquots in the same RNeasy spin column. Discard the flow-through after each centrifugation.
- Add 700 µl Buffer RW1 to the RNeasy spin column.
- Close the lid gently, and centrifuge for 30 s at 15,000 rcf to wash the spin column membrane. Discard the flow-through.
- Add 500 µl Buffer RPE to the RNeasy spin column.
- Close the lid gently, and centrifuge for 30 s at 15,000 rcf to wash the spin column membrane. Discard the flow-through.
- Add 500 µl Buffer RPE to the RNeasy spin column.
- Close the lid gently, and centrifuge for 1 min at 15,000 rcf to wash + dry the spin column membrane.
- Place the RNeasy spin column in a new 2 ml collection tube (supplied), and discard the old collection tube with the flow-through.
- Close the lid gently, and centrifuge at full speed for 2 min.
- Place the RNeasy spin column in a new 1.5 ml collection tube (supplied)
- Add 50 µl RNAse-free water directly to the spin column membrane. Close the lid gently, and centrifuge for 1 min at 12,000 rcf to elute the RNA.
- If higher concentration is desired, re-apply this eluate to the filter membrane and repeat the centrifuge step.
3/16/26: All final blocks embedded, details in google sheet and notebook
https://docs.google.com/spreadsheets/d/1y8tZUTPmXJmq1QOfvcBU3mhVc6VYH7Xe-fA4zXSnTZ4/edit?usp=sharing
Embedded blocks were sectioned 3/17 and 3/19:
| Block_Label | Embedding notes - how to section | Block_Description | Section_Date | Section_Notes | Block_Status | Slide_1_label | Slide_1_Status | Slide_2_label | Slide_2_Status | Slide_3_label | Slide_3_Status | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| POC_1-A | discs | 48 hr OCT:30% test | 20260317 | Second slide is better - looking pretty good | in -80 | 0317_02 | H&E Stained 3/20 | 0317_03 | H&E Stained 3/20 | ||||||
| POC_2-A | discs | 48 hr | 20260317 | Tore a lot, one branch I think | in -80 | 0317_04 | H&E Stained 3/20 | 0317_05 | H&E Stained 3/20 | ||||||
| POC_3 | one disc | 72hr, no sucrose | 20260317 | Sectioned super well | in -80 | 0319_10 | H&E Stained 3/20 | 0319_11 | H&E Stained 3/20 | 0319_12 | H&E Stained 3/20 | ||||
| POC_4 | one disc | 72 hr OCT:30% test | 20260317 | Sectioned super well; collected 3 tubes, use #3 | in -80 | 0317_08 | H&E Stained 3/20 | 0317_09 | H&E Stained 3/20 | 0319_09 | H&E Stained 3/20 |
| Block_Label | Embedding notes - how to section | Block_Description | Section_Date | Section_Notes | Block_Status | Slide_1_label | Slide_1_Status | Slide_2_label | Slide_2_Status | Slide_3_label | Slide_3_Status |
|---|---|---|---|---|---|---|---|---|---|---|---|
| POR_1 | section 90 degrees | 72hr, no sucrose | 20260319 | went ok; got automatic sectioning to work | in -80 | 0319_08 | H&E Stained 3/20 | ||||
| POR_2 | section 90 degrees | 72 hr OCT:30% test | 20260319 | tore a little but pretty good | in -80 | 0319_06 | in -80 | 0319_07 | H&E Stained 3/20 | ||
| POR_3 | section 90 degrees | 96 hr OCT:30% test | 20260319 | 2nd slide has one section at 20 um | in -80 | 0319_03 | H&E Stained 3/20 | 0319_04 | H&E Stained 3/20 | ||
| POR_4 | section 90 degrees | 96 hr OCT:30% test, expect rip | 20260319 | took about an hour to get in the groove, but when it did it worked great | in -80 | 0319_01 | H&E Stained 3/20 | 0319_02 | H&E Stained 3/20 | ||
| POR_5 | section 90 degrees | 96 hr | 20260319 | tore a lot, tissue coming out of OCT | in -80 | 0319_05 | in -80 |
| Block_Label | Embedding notes - how to section | Block_Description | Section_Date | Section_Notes | Block_Status | Slide_1_label | Slide_1_Status | Slide_2_label | Slide_2_Status | Slide_3_label | Slide_3_Status | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MON_1 | section like normal | 48 hr OCT:30% test - had some skeleton, expect rippage | 20260319 | 2nd tube is better | in -80 | 0319_10 | H&E Stained 3/20 | 0319_11 | H&E Stained 3/20 | 0319_12 | H&E Stained 3/20 | |||||
| MON_2 | section 90 degrees | 48 hr OCT:30% test | 20260317;20260319 | Hard to section :/ Ripped a lot. sectioned second slide at 20 um. Tried again 3/19. coming out of OCT a lot | in -80 | 0317_08 | H&E Stained 3/20 | 0317_09 | H&E Stained 3/20 | 0319_09 | H&E Stained 3/20 | |||||
| MON_3 | section 90 degrees | 48 hr | 20260317 | Hard to section :/ Ripped a lot because it was so far to the bottom of the mold. sectioned at 20 um | in -80 | 0317_10 | H&E Stained 3/20 | |||||||||
| MON_4 | section 90 degrees | 48 hr, expected to be messed up though | 20260319 | Coming out of OCT too much. 1 tube but no slides | in -80 | In -80 | In -80 | H&E Stained 3/2 | H&E Stained 3/2 |
3/25/26: Tissue scrolls extraction
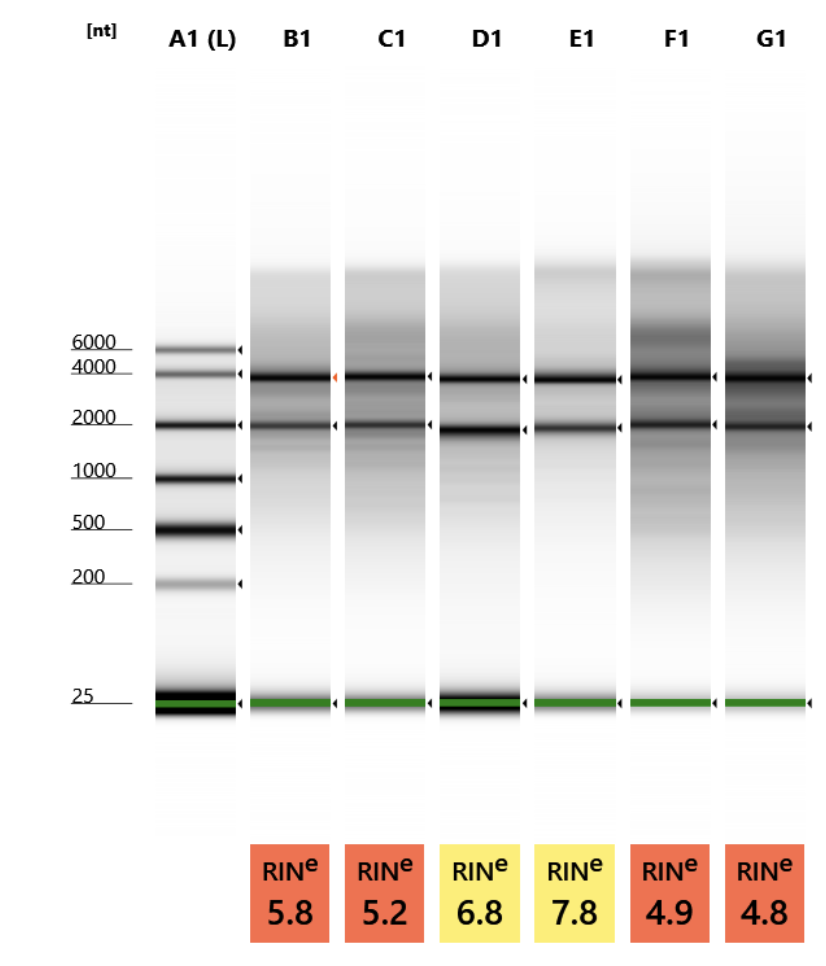
| sample_id | concentration | RIN |
|---|---|---|
| Scroll - POC_3 (EDTA 72hr, no sucrose) | 2.47 ng/uL | 5.8 |
| Scroll - POC_4 (EDTA 72hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 3.24 ng/uL | 5.2 |
| Scroll - POR_2 (EDTA 72hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 1.13 ng/uL | 6.8 |
| Scroll - POR_3 (EDTA 96hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 2.18 ng/uL | 7.8 |
| Scroll - MON_2 (EDTA 48hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 8.22 ng/uL | 4.9 |
| Scroll - MON_3 (EDTA 48hr, Sucrose w/o extra 30% Sucrose:OCT incubation) | 7.77 ng/uL | 4.8 |
Full results can be found here
| sample_id | concentration | RIN | example pic | slidescan | morphology assessment/notes |
|---|---|---|---|---|---|
| Scroll - POC_3 (EDTA 72hr, no sucrose) | 2.47 ng/uL | 5.8 |  |
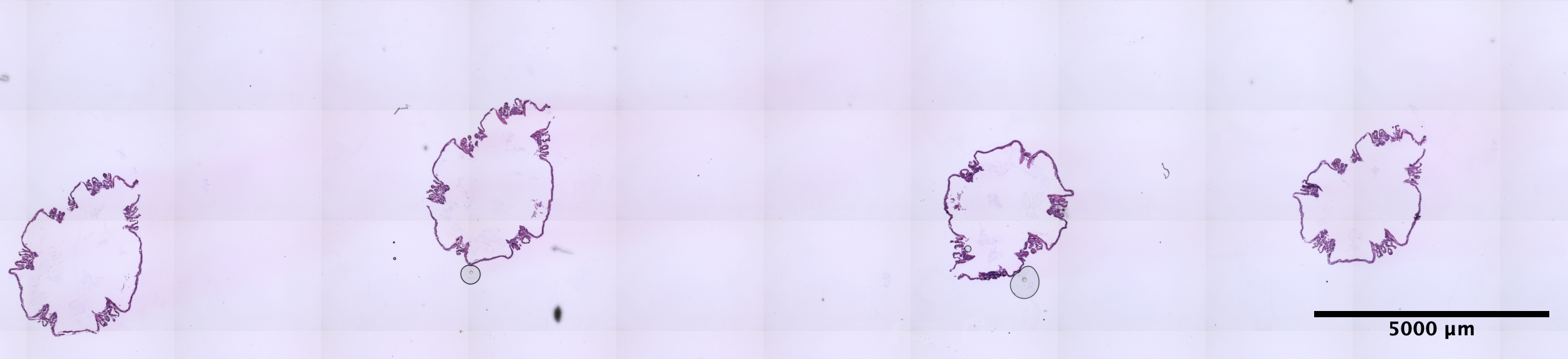 |
calcioblast morphplogy not preserved as well as POC_4 |
| Scroll - POC_4 (EDTA 72hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 3.24 ng/uL | 5.2 |  |
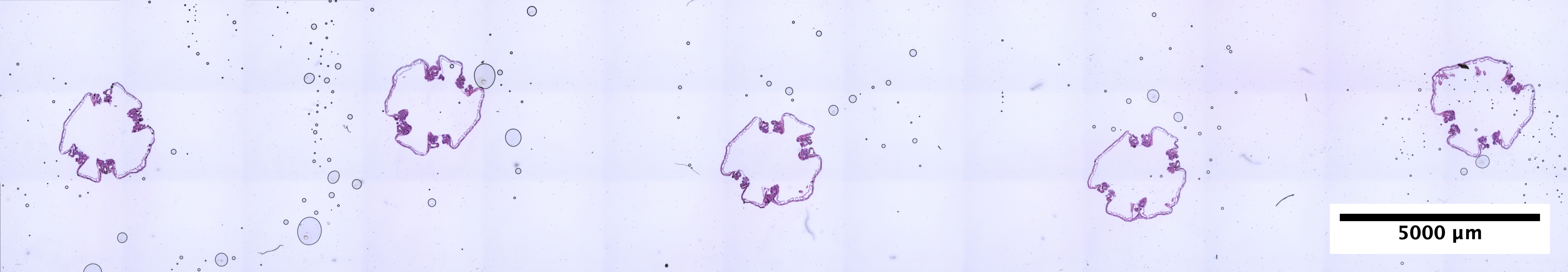 |
looks really really great |
| Scroll - POR_2 (EDTA 72hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 1.13 ng/uL | 6.8 | 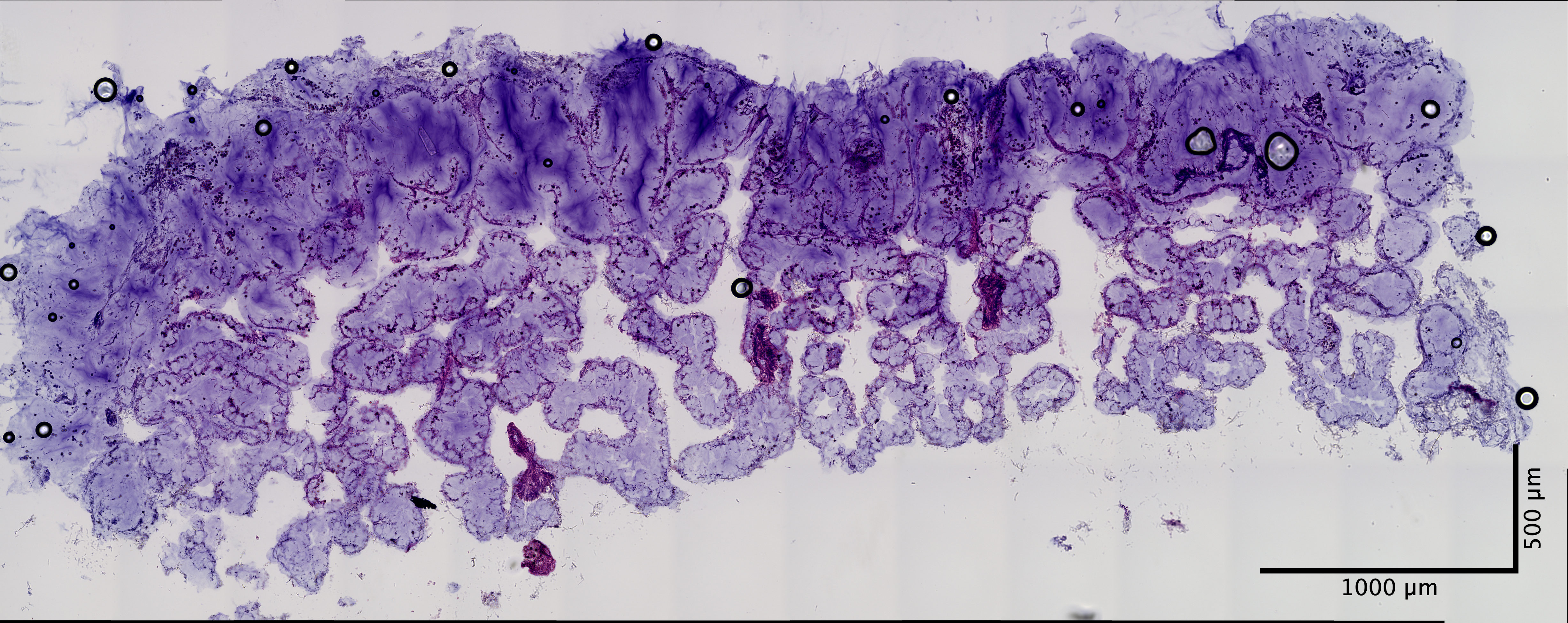 |
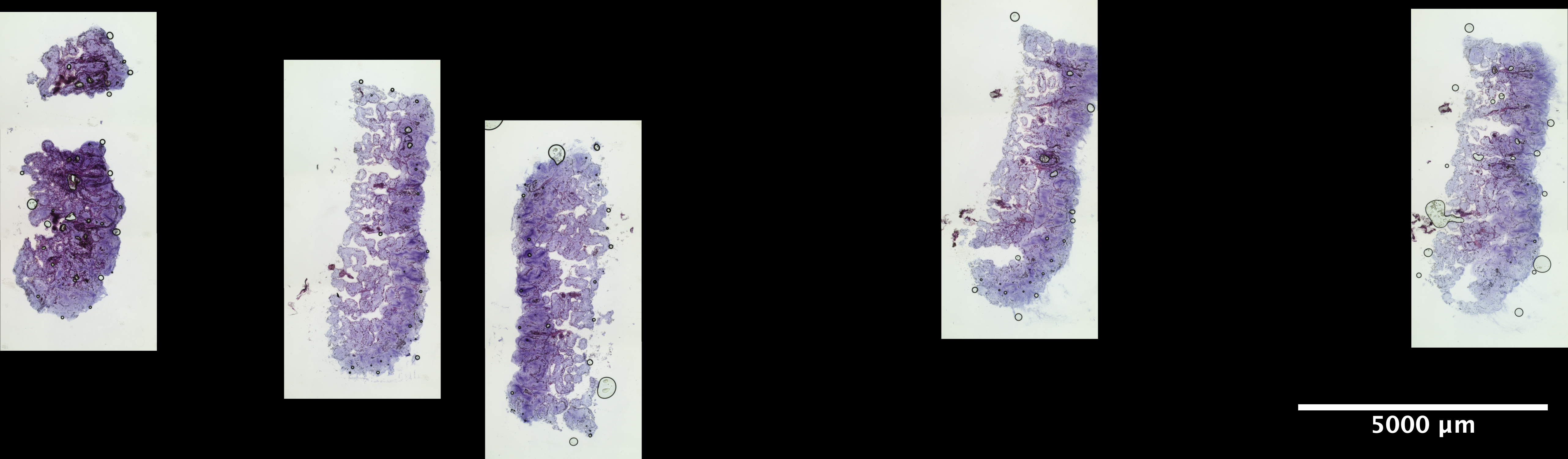 |
looks great |
| Scroll - POR_3 (EDTA 96hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 2.18 ng/uL | 7.8 |  |
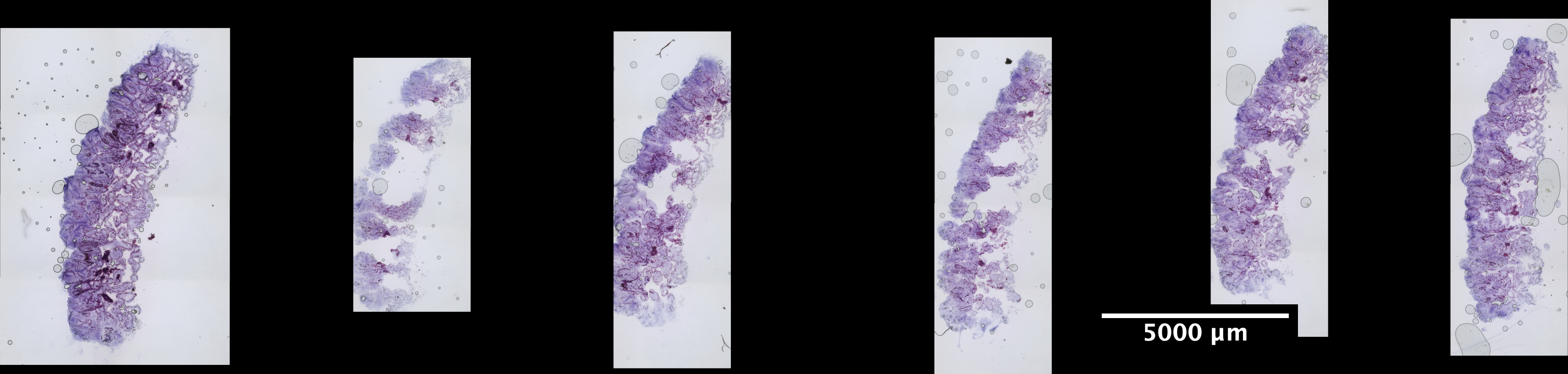 |
looks really really great |
| Scroll - MON_2 (EDTA 48hr, Sucrose w/ extra 30% Sucrose:OCT incubation) | 8.22 ng/uL | 4.9 |  |
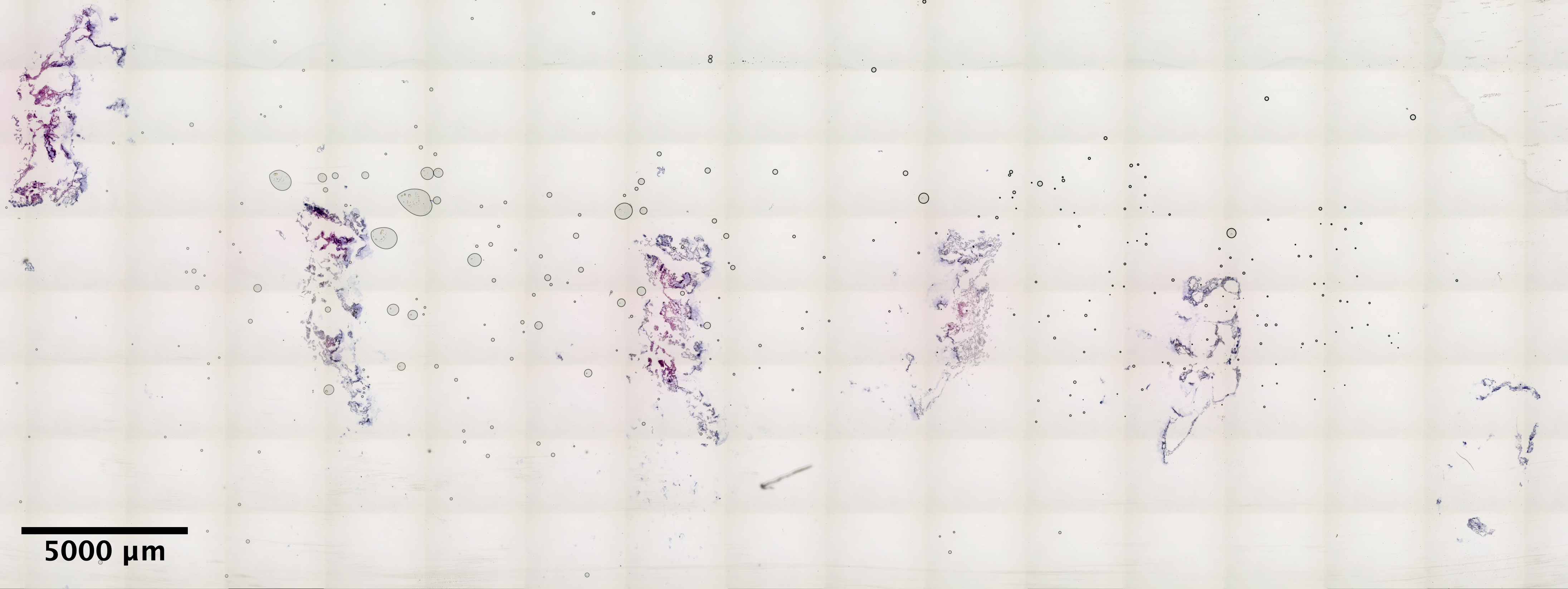 |
really jumbled tissue, hard to tell what is what |
| Scroll - MON_3 (EDTA 48hr, Sucrose w/o extra 30% Sucrose:OCT incubation) | 7.77 ng/uL | 4.8 |  |
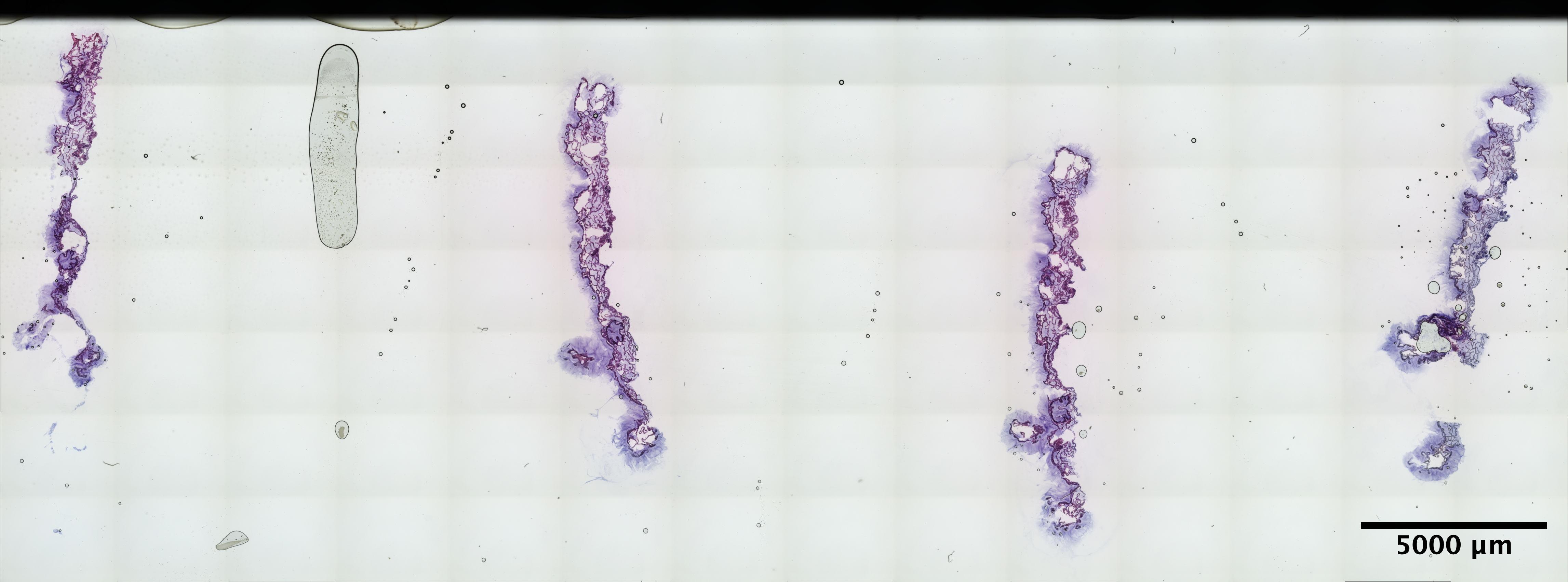 |
still not stellar, but maybe the best MON sections we’ve ever gotten |
Written on March 10, 2026